You are here: vision-research.eu » Vision Research » The Young Researchers View » Martial Mbefo (Q04-2019)
The Research of Martial Mbefo
 |
Background and research
I moved to Prof Arsenijevic’s lab at Jules Gonin’s Eye Hospital in Lausanne after having completed my PhD degree in neurosciences at the Swiss Federal Institute of Technology (EPFL, Switzerland). My PhD project focused on understanding the role of A-syn posttranslational modifications during the physiopathology of Parkinson Disease. Moving to Yvan’s lab has been fun, as it provided me with the unique opportunity to explore the mechanistic aspects of neuroretina cell degeneration and sight loss in retinitis pigmentosa (RP).
Results from independent research groups exploring the inherited retinal degeneration have proven a crucial role for gene regulation changes in photoreceptor cell death and rescue. A deep analysis of certain relevant pathways such DNA methylation [1], HDAC activation [2], could suggest a role for epigenetic dysfunction in RD and related diseases.
To this end, my current research at the unit of Retinal Degeneration and Regeneration focuses on developing a bunch of epigenetic tools in order to understand whether abnormal histone posttranslational modifications and microRNA expressions occur in RD, and how they can promote or modulate the expression of a given gene associated with disease.
Role of Histones modifications in Retinal Degeneration
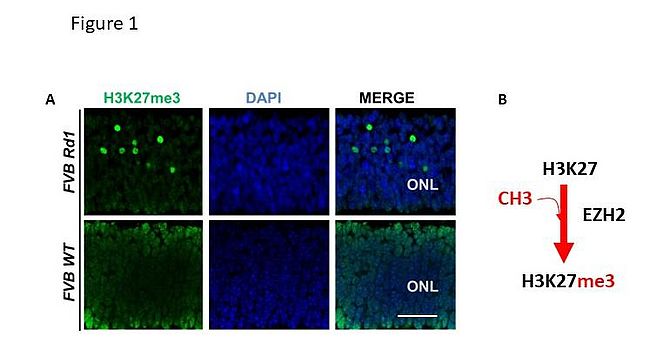
In views of the beneficial role that the Bmi1 deficiency generates to the Rd1 mice [3] and given that BMI1 is member of the Polycomb group ring finger protein (PcG) which modifies the chromatin through the modifications of histone tails to repress gene activity [4], we asked whether disruption of gene regulatory network in Rd1 could be epigenetic dependent. We monitored the expression pattern of key histones marks that contribute to repressive (H3K27me3, H3K9me2) and permissive (H3K4me3) chromatin in Rd1 and WT mice. We found that the H3K27me3 mark was robustly hypermethylated in the retina of the Rd1 mice at onset and during the process of cell death (Fig1, poster ARVO 2015), and was further confirmed by independent research group [5].
The methylation of H3K27me3 is often mediated by the Enhancer of Zeste Homolog 2 (EZH2) in vertebrates and mammals [6] and its altered expression promotes premature retinal differentiation and contributes to retinal degeneration in mice [7]. Consistent with the hypermethylation found in Rd1, EZH2 expression was enhanced in purified photoreceptor extracts obtained from Crx-Gfp Rd1 compared to Crx-Gfp WT at p12 as revealed by Western blot.
In addition to our observation that the hyper trimethylation of H3K27me3 occurs upstream CDK4 and the TUNEL positive cells at p12 and does not colocalizes with the cGMP which is the early marker of cell death in Rd1, suggesting that it is an early event. Current investigations rely on the analysis of CHIP-Seq and mRNA-Seq datasets. In our analysis workflow we are developing efficient bioinformatic tools enabling to establish the correlation between histones marks depositions at various gene regulatory regions and their expression pattern. Relevant gene candidates with differential histone depositions were selected and are currently being validated in vivo.
MicroRNA and gene regulation in RD
MiroRNAs are part of the epigenetic regulatory mechanisms that inhibit gene expression by targeting mRNA. In stress conditions, a combined activity of BMI1 and EZH2 can affect the pool of microRNAs by promoting differential histone marks deposition at their promoter regions. Although a set of microRNA families was identified to actively participate to the process of neurodegenerative disorders, including PD, AD and HD [8], very little is known about microRNAs during retinal degeneration (RD). Because a given microRNA may have different targets, their dysregulation may have a great impact at disease onset, and during its evolution.
We identified 42 microRNAs that were significantly (FDR <0.05, Foldchange ±2) differentially expressed in Rd1 microarray. Three microRNAs were among the miRNAs that were downregulated the most. They were shown to either play a role on photoreceptor differentiation or to be altered in tumor cells [9, 10].
Our preliminary studies on this microRNAs suggest a potential role on cell cycle repression as revealed by validated targets gene list obtained from miRTarBase database. Preliminary CHIP-seq analysis of the H3K27m3 mark, revealed that the transcription start site of most downregulated microRNA was enriched of this mark in Rd1 retina. Target predictions analysis of downregulated miRNAs and STRING protein interaction network analysis allowed to identify several hubs that were in accordance with some disease features already described and observed with our mRNA-Seq data.
As such, these data suggest that alteration of microRNAs biogenesis in Rd1 mice occurs in EZH2 dependent manner and is associated with the disease progression.
Significance of H3K27me3 marks in retinal degeneration.
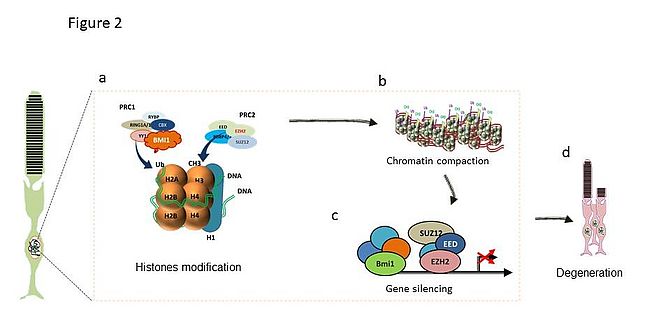
We believe that a dysregulation of histones marks H3K27me3 may occur during the degenerative stage in response to the Pde6b deficiency in Rd1 mice (Fig2). Histones binding to promoters regions of genes essential for photoreceptor survival can be dramatically hypermethylated, thereby promoting their unexpected repression and subsequent degeneration.
The H3K27me3 regulates gene expression either in coordination with H3K9me3 (permanently inactive promoter), with H3K4me3 (bivalent or poised chromatin promoters), or alone as temporarily inactive or repressive chromatin. These features and mostly the temporarily inactive state upon H3K27me3 open insights for controlling the H3K27me3 landscape in health and disease through a wide range of existing and newly developed pharmacological inhibitors. A deep understanding of how H3K27me3 is regulated, for example by controlling the EZH2 activity in retinal disorders may be crucial for understanding the key mechanisms controlling retinal degeneration.
References
- Farinelli, P., et al., DNA methylation and differential gene regulation in photoreceptor cell death. Cell Death Dis, 2014. 5: p. e1558.
- Yuan, H., et al., Involvement of HDAC6 in ischaemia and reperfusion-induced rat retinal injury. BMC Ophthalmol, 2018. 18(1): p. 300.
- Zencak, D., et al., Retinal degeneration depends on Bmi1 function and reactivation of cell cycle proteins. Proc Natl Acad Sci U S A, 2013. 110(7): p. E593-601.
- Abdouh, M., et al., The Polycomb Repressive Complex 1 Protein BMI1 Is Required for Constitutive Heterochromatin Formation and Silencing in Mammalian Somatic Cells. J Biol Chem, 2016. 291(1): p. 182-97.
- Zheng, S., et al., DZNep inhibits H3K27me3 deposition and delays retinal degeneration in the rd1 mice. Cell Death Dis, 2018. 9(3): p. 310.
- Margueron, R. and D. Reinberg, The Polycomb complex PRC2 and its mark in life. Nature, 2011. 469(7330): p. 343-9.
- Yan, N., et al., Postnatal onset of retinal degeneration by loss of embryonic Ezh2 repression of Six1. Sci Rep, 2016. 6: p. 33887.
- Shah, P., et al., MicroRNA Biomarkers in Neurodegenerative Diseases and Emerging Nano-Sensors Technology. J Mov Disord, 2017. 10(1): p. 18-28.
- McEvoy, J., et al., Analysis of MDM2 and MDM4 single nucleotide polymorphisms, mRNA splicing and protein expression in retinoblastoma. PLoS One, 2012. 7(8): p. e42739.
- Montoya, V., et al., Novel miRNA-31 and miRNA-200a-Mediated Regulation of Retinoblastoma Proliferation. PLoS One, 2015. 10(9): p. e0138366.

Martial Mbefo, PhD.
Research Scientist
Jules-Gonin Eye Hospital,
Oculogenetic and Gene Therapy,
University of Lausanne,
Avenue de france 15
1004 Lausanne,
Switzerland
Email: martial.mbefo[at]fa2.ch