You are here: vision-research.eu » Vision Research » The Young Researchers View » Kinga Bujakowska (Q02-2012)
Searching for the underlying genetic causes of retinal degenerations
 |
Inherited retinal disorders (IRD) are an important cause of blindness and severe impairment of vision worldwide. IRDs are a very heterogeneous group of diseases ranging from blindness at birth or within the first year of life, as in Leber Congenital Amaurosis (LCA), to milder forms, such as congenital stationary night blindness (CSNB). So far, mutations in approximately 250 genes or gene loci have been associated with various forms of retinal degenerations; however we still do not know the molecular cause of the disease in a large proportion of patients. One of my research interests is to find these new genetic defects leading to IRD.
Working in the Insitut de la Vision in the team of Dr Isabelle Audo and Dr Christina Zeitz, I have the opportunity to be involved in investigating recessive cases of retinitis pigmentosa (RP) and congenital stationary night blindness (CSNB). DNA samples of patients with a precise diagnosis are screened for genetic defects using a combination of different methods. In the first instance, known mutations were in many cases excluded with ASPER mutation detection array or by Sanger sequencing of prevalent genes. To find novel genes involved in IRD we primarily use four methods: 1) homozygosity mapping, 2) candidate gene approach, 3) linkage analysis, 4), next generation sequencing.
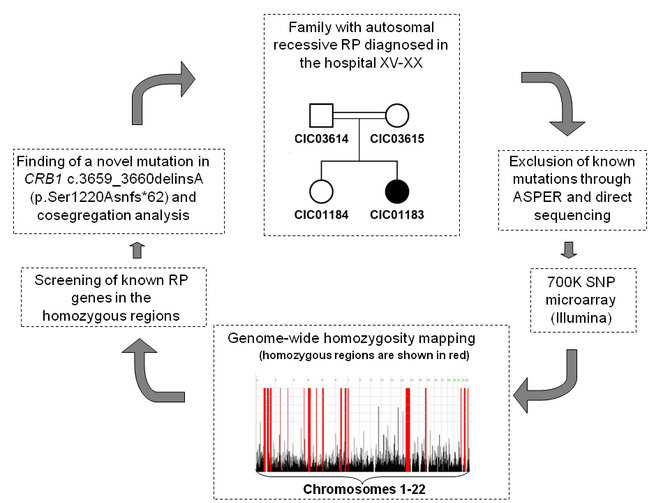
We use homozygosity mapping for small consanguineous families and have, for example, successfully applied the technique to one family carrying a CRB1 mutation (Figure 1). After initial exclusion of known mutations with ASPER mutation detection array, the affected and unaffected family members were analysed with a dense SNP microarray (Illumina HumanOmniExpress). As the family is consanguineous, we expected the disease causing mutation to be found in homozygous regions (marked in red on figure 1). Screening of known RP genes in these regions revealed a frameshift mutation c.3659_3660delinsA in CRB1, leading to protein truncation p.Ser1220Asnfs*62 (Bujakowksa et al. 2012). Similarly we found novel mutations in TULP1, PDE6B and DFNB31 (Audo et al. 2010).
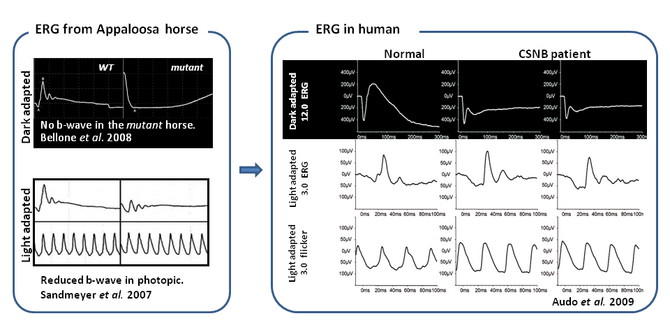
One way of discovering new genetic defects implicated in IRD is by performing candidate gene approaches. This strategy consists of a literature search of animal models with phenotypes resembling those of human patients. In this way our group and others have proven that mutations in TRPM1 in humans lead to CSNB (Audo et al 2009). Screening of this gene was undertaken after the publication of an article which stated that the differential expression of TRPM1 in Appaloosa horses is the potential cause of CSNB (Bellone et al 2008).
Recently we have applied next generation sequencing to find novel genetic defects underlying retinal disorders. With this technique we have sequenced whole exomes of patients and selected unaffected family members and were able to find mutations in a new gene implicated in CSNB, G protein-coupled receptor 179 (GPR179) (Audo et al. 2012). Genetics is a fast developing field and the whole-genome approaches such as next-generation sequencing or dense SNP arrays have greatly accelerated the discovery of molecular causes of inherited diseases. The important and exciting discovery of new genes implicated in retinal disorders facilitates understanding of pathologies, enables the development of future therapies and contributes to the understanding of retinal physiology and cellular pathways.
Selected Publications
- Audo I, Bujakowska K, Orhan E, Poloschek CM, Defoort-Dhellemmes S, Drumare I, Kohl S, Luu TD, Lecompte O, Zrenner E, Lancelot ME, Antonio A, Germain A, Michiels C, Audier C, Letexier M, Saraiva JP, Leroy BP, Munier FL, Mohand-Saïd S, Lorenz B, Friedburg C, Preising M, Kellner U, Renner AB, Moskova-Doumanova V, Berger W, Wissinger B, Hamel CP, Schorderet DF, De Baere E, Sharon D, Banin E, Jacobson SG, Bonneau D, Zanlonghi X, Le Meur G, Casteels I, Koenekoop R, Long VW, Meire F, Prescott K, de Ravel T, Simmons I, Nguyen H, Dollfus H, Poch O, Léveillard T, Nguyen-Ba-Charvet K, Sahel JA, Bhattacharya SS, Zeitz C. Whole-Exome Sequencing Identifies Mutations in GPR179 Leading to Autosomal-Recessive Complete Congenital Stationary Night Blindness. Am J Hum Genet. 2012 Feb 10;90(2):321-30.
- Audo I, Bujakowska KM, Leveillard T, Mohand-Said S, Lancelot ME, Germain A, Antonio A, Michiels C, Saraiva JP, Letexier M, Sahel JA, Bhattacharya SS, Zeitz C. Development and application of a next-generation-sequencing (NGS) approach to detect known and novel gene defects underlying retinal diseases. Orphanet J Rare Dis. 2012 Jan 25;7(1):8.
- Bujakowska K, Audo I, Mohand-Saïd S, Lancelot ME, Antonio A, Germain A, Léveillard T, Letexier M, Saraiva JP, Lonjou C, Carpentier W, Sahel JA, Bhattacharya SS, Zeitz C. CRB1 mutations in inherited retinal dystrophies. Hum Mutat. 2012 Feb;33(2):306-15. doi: 10.1002/humu.21653. Epub 2011 Dec 27.
- Audo I, Bujakowska K, Mohand-Saïd S, Tronche S, Lancelot ME, Antonio A, Germain A, Lonjou C, Carpentier W, Sahel JA, Bhattacharya S, Zeitz C. A novel DFNB31 mutation associated with Usher type 2 syndrome showing variable degrees of auditory loss in a consanguineous Portuguese family. Mol Vis. 2011;17:1598-606. Epub 2011 Jun 15.
- Audo I, Bujakowska K, Mohand-Saïd S, Lancelot ME, Moskova-Doumanova V, Waseem NH, Antonio A, Sahel JA, Bhattacharya SS, Zeitz C. Prevalence and novelty of PRPF31 mutations in French autosomal dominant rod-cone dystrophy patients and a review of published reports. BMC Med Genet. 2010 Oct 12;11:145. Review.
- Audo I, Kohl S, Leroy BP, Munier FL, Guillonneau X, Mohand-Saïd S, Bujakowska K, Nandrot EF, Lorenz B, Preising M, Kellner U, Renner AB, Bernd A, Antonio A, Moskova-Doumanova V, Lancelot ME, Poloschek CM, Drumare I, Defoort-Dhellemmes S, Wissinger B, Léveillard T, Hamel CP, Schorderet DF, De Baere E, Berger W, Jacobson SG, Zrenner E, Sahel JA, Bhattacharya SS, Zeitz C. TRPM1 is mutated in patients with autosomal-recessive complete congenital stationary night blindness. Am J Hum Genet. 2009 Nov;85(5):720-9.
Institut de la Vision
Inserm UMR S968
17, Rue Moreau
75012 Paris
France
Tel.: +33 1 53 46 25 40
FAX: +33 1 53 46 26 02